ContentslistsavailableatScienceDirect
Journal
of
Infection
and
Public
Health
j o u r n al ho me p ag e :h t t p : / / w w w . e l s e v i e r . c o m / l oc a t e / j i p h
Hepatitis
E
in
Italy:
A
silent
presence
Carlo
Mauceri
a,
Maria
Grazia
Clemente
a,
Paolo
Castiglia
b,
Roberto
Antonucci
a,
Kathleen
B.
Schwarz
c,∗aPediatricClinic,DepartmentofSurgical,MicrosurgicalandMedicalSciences,UniversityofSassariMedicalSchool,07100Sassari,Italy bDepartmentofBiomedicalSciences—HygieneandPreventiveMedicineUnit,University-AOUofSassari,07100Sassari,Italy cPediatricLiverCenter,JohnsHopkinsUniversitySchoolofMedicine,Baltimore21287,MD,USA
a
r
t
i
c
l
e
i
n
f
o
Articlehistory:
Received3February2017 Receivedinrevisedform3July2017 Accepted4August2017 Keywords: HepatitisEinfection Epidemiology Seroprevalence Riskfactors Immigrants Italy
a
b
s
t
r
a
c
t
HepatitisEvirus(HEV)wasdiscoveredinthe1980sandhasbeenconsideredasbeingconfinedto devel-opingcountries.ThepurposeofthiscriticalreviewwastodeterminethereportedHEVseroprevalence ratesinItaly,toidentifypredisposingfactorsandindividualsatriskandtoassesspossibleimportation ofHEVbyimmigrants.Acriticalreviewof159articlespublishedinPubMedfrom1994todatewas done.Only27originalreportsof50ormoresubjects,writtenintheEnglishorItalianlanguage,were included.Overthreedecades,theHEVseroprevalencevariedfrom0.12%to49%,withthehighestrates beingreportedfromthecentralregionofItaly.Riskfactorsincludedingestionofrawporkorpotentially contaminatedfood.Theseroprevalenceamongimmigrantsrangedfrom15.3%to19.7%inApulia.Italyhas apopulationof60656000;thetotalnumberofindividualssurveyedwasonly21.882(0.036%).Anational epidemiologicalsurveyprogramisneededtocapturemorecomprehensiveseroprevalencedata.
©2017 TheAuthors.PublishedbyElsevierLimitedonbehalfofKingSaudBinAbdulazizUniversity forHealthSciences.ThisisanopenaccessarticleundertheCCBY-NC-NDlicense(http:// creativecommons.org/licenses/by-nc-nd/4.0/). Contents Introduction...1 Methods...2 Results...2 Discussion...4 Conclusion...6 Funding ... 6 Competinginterests...6 Ethicalapproval...6 Acknowledgments...6 References...6
Abbreviation: Assay-1,Dia.Pro;Assay-2, Abbott;Assay-3,Wantai;Assay-4, Adaltis;HEV,HepatitisEvirus.
∗ Correspondingauthorat:JohnsHopkinsUniversitySchoolofMedicine, Pedi-atricLiverCenter,CMSC2-116,600NorthWolfeSt.Baltimore,Md.21287,USA. Fax:+14109551464.
E-mailaddresses:[email protected](C.Mauceri),
[email protected](M.GraziaClemente),[email protected](P.Castiglia),
[email protected](R.Antonucci),[email protected](K.B.Schwarz).
Introduction
HepatitisEvirus(HEV)istheubiquitousetiologicalagentof entericnon-Aviralhepatitisand itrepresentsanongoing inter-nationallychallenging issue for publichealth. Annually, HEVis responsiblefor3.3millionnewsymptomaticinfectionswithfatal outcomesin 56600individuals worldwide [1,2].Three decades afteritsdiscoveryduringanoutbreakofunexplainedhepatitisin Afghanistan[3],notonlyisitsoriginunknownbutthemodesof transmissionremainfarfrombeingclearlyunderstoodinthe indus-trializedworld.HEVisasmallhepatotropicsingle-strandedRNA virus,thesolememberoftheHepeviridaefamily,belongingtothe https://doi.org/10.1016/j.jiph.2017.08.004
1876-0341/©2017TheAuthors.PublishedbyElsevierLimitedonbehalfofKingSaudBinAbdulazizUniversityforHealthSciences.Thisisanopenaccessarticleunderthe CCBY-NC-NDlicense(http://creativecommons.org/licenses/by-nc-nd/4.0/).
viousgenomicsequenceanalysishadrevealedtheexistenceoffour well-definedmammaliangenotypesandatleast24sub-genotypes, withaspecificgeographicdistribution[6].Indeed,the epidemi-ologyandpathogenicityofHEVobservesabimodalpatternthat differsinemergingnationsandoccidentalcountries.Genotypes1 and2causelargeoutbreaksandepidemicsmainlyamongyoung adultsinAfrica, CentralandSouthernAsia,CentralAmerica and theMiddleEast[7–12].Theinfectionisacquiredpredominately throughafecal–oralrouteanditisassociatedwithanunusually highmortalityrateduringpregnancy[13].Poorsanitationand con-taminatedwater sourcesareprecipitatingfactors.Anincreasing numberofsporadicandsmalllocallyacquiredoutbreakshavebeen reportedinNorthernAmerica,Australia,Europe,ChinaandJapan [14–19].Genotypes3and4arelessvirulentstrainsidentifiedasthe causativeagentsofsubclinicalandclinicalinfectionintheelderly population.HEVismostlytransmittedzoonoticallywiththe inges-tionofrawandundercookedfood.Todata,eightgenotypeshave beendetectedand4ofwhichareconfinedtoanimalspecies: geno-type5and6inJapanesewildboar(Scrofascrofaleucomystax)[20] andgenotype7and8respectivelyindromedarycamels(Camelus dromedaries)andBactriancamels(Camelusbactrianus)[21].
InEurope,HEVseroprevalenceestimatesrangedin the gen-eralpopulationfrom7.5% to31.9% withtheaverageratebeing 19.16%;ratesincreasewithage[22].However,therealprevalence couldbeunderestimatedduetothedifferenceintestsensitivity andthefrequentlyasymptomaticcourseofthedisease.Overall, itislikelythatcurrentgeopoliticalinstabilityandtheconsequent massiveimmigrationwouldleadtowardsthelocalintroductionof newpathogenicvariantsandmodifytheknownepidemiologyin Westerncountries.
Methods
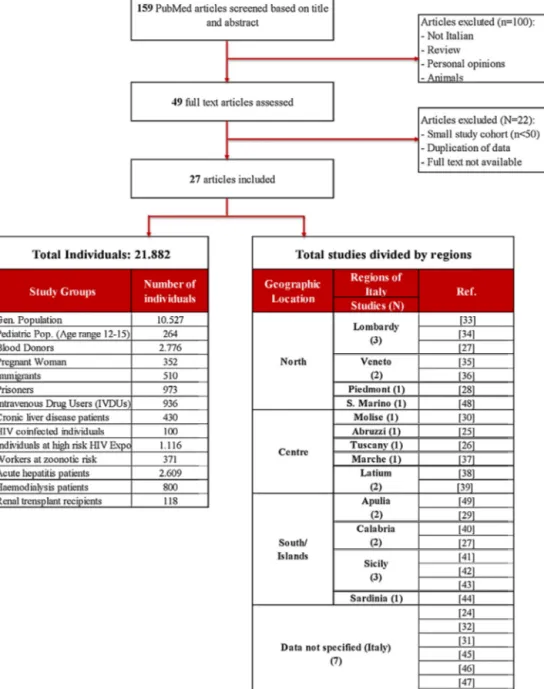
ThecriticalreviewisbasedonaliteraturesearchonPubMed, usingthekeywords“hepatitisEinItaly”and“hepatitisE seropreva-lenceinItaly”.StudiespublishedfromJanuary1994andMay2017 wereincludedaccordingtothefollowingcriteria:studiesprovided clearinformationregardingtheseroprevalencerateattheregional ornationallevelandincludedatleast50samplesinthecohort (Fig.1).Noagerestrictionwasobservedandallstudieswere writ-tenintheEnglishorItalianlanguage.Thestatisticalanalysesofthe reportedregionalseroprevalenceshavebeendone.Weusedthe screeningmethodstoadjusttheprevalencevalueaccordingtothe sensitivityandspecificityoftheassay.Onlyestimated seropreva-lencerateswithalowerpositivevalueforC.I.at95%havebeen consideratestatisticallysignificant.
Accordingtothedataavailable,wefocusedonfourteenstudy cohorts: general population, blood donors, pregnant women, thepediatric population, acute hepatitis patients, chronic liver disease patients, hemodialysis patients, immigrants, prisoners, intravenousdrugusers,HIVco-infectedindividuals,HIV-exposed and/orinfectedindividualsandworkerswithcontactwith poten-tial zoonotic reservoirs (abattoir workers, laboratory workers exposedtobiologicalswinematerial,animalbreeders, veterinari-ansandfarmers)andrecipientsofrenaltransplants.Studiesthat didnot meet the above-mentionedcriteria, provided duplicate data,personal opinion or international reviews,were excluded fromthecriticalreview.
Results
159publicationswereidentifiedbytitleandabstractthrough aPubMedsearchand27articleswereincludedinthefinaldata
and/oranti-HEVIgM,representingonly0,036%ofthecurrentItalian population[23].Theseroprevalenceratesrangedfrom0,12%to49% amongthestudycohort[24,25].TheAbruzziregionwasfoundtobe ahyper-endemicregionwithaseroprevalencerateof49%among blooddonors[25].Aseroprevalencestudyof132blooddonor resi-dentsinTuscanyhasreportedratesof9.1%[26],whichissimilarto the9%rateofthesamecohortofindividualsintheLatiumregionin 2009(furtherdatanotshown).[25].Aseroprevalenceof9%wasalso reportedinthegeneralpopulationofAbbiategrasso,inLombardy; thehighestassessedamongthenorthernregionsofItaly[27]. Con-versely,thelowestseroprevalenceof1.3%and2.7%wasreported inPiedmontandApuliaamongtheopenpopulation,respectively inthenorthandsouthofItaly[28,29].Thehighestrateamongthe southernregionswasreportedinCalabria(Casanova)witha sero-prevalenceof17.8%[27].Basedonthedataonagereportedby17 ofthe27studies,wecalculatedameanageof42.28yearsforthe HEVpositivesubjects.Moreover,59,26%weremales,accordingto theinformationprovidedby21studies.Overall,theseroprevalence increasedinassociationwithageandnorelevantvariationrelated togenderhasemergedfromanystudy.However,onlyonepediatric studywithaprevalenceof0.4%wasfoundinMolise[30].
Thestudyincluded:10.527individualsfromthegeneral pop-ulation cohort, 2776 blood donors, 352 pregnant woman, 264 individualsatpediatricage,2.609patientsaffectedbyacute hepati-tis,800individualsinhemodialysis,118renaltransplantrecipients, 430chronicliverdiseasepatients,371atzoonoticriskworkers,510 immigrantsand3.125athigh-riskindividuals(100HIVinfected and1116withsexualoroccupationalexposure,936intravenous drugusersand973prisoners).
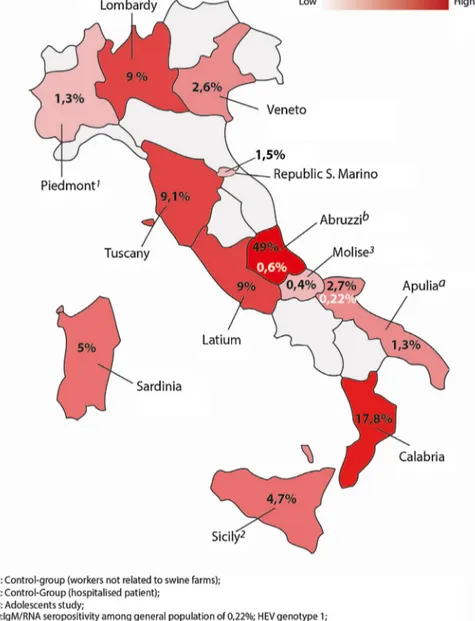
The selected articleswere from 13 different regional areas: Abruzzi(n=1),Apulia(n=2),Calabria(n=2),Latium(n=2), Lom-bardy(n=3),Marche(n=1),Molise(n=1),Molise(n=1),Piedmont (1),Sardinia(n=1),Sicily(n=3),Republicof SanMarino (n=1), Tuscany(n=1),andVeneto(n=2)(Fig.2).
Theremaining7studiesprovidedinformationatanationallevel only.Overthreedecades,169casesofhepatitisEwerelinkedto travel inhighendemic countries. Nearly90% of them occurred intravelers returning fromBangladesh, Indiaand Pakistan.The remainingcaseswerediagnosedinpatientswhotraveledinAngola, Somalia,MoroccoandGreen Cape.Secondary andintra-familiar infectionhasbeendescribedbytwo studieswitharateof2.6% and4.5%respectively[31,32].Generally,ahigherprobabilitytobe positiveforanti-HEVantibodieshasbeenassociatedwith immi-grantsby thedifferentstudies focusingon socio-economicand demographicvariables(healthypopulation,prisonersandat-risk categories).Genotype1(G1),subtype1aand1c,hasbeenisolated inallimportedcasesofHEV.Genotype3(G3),subtype3e,3fand3h, hasbeenassociatedwithalocalsourceofinfection.Nosignificant differencesintheclinicalcourseofthediseasecausedbytheG1 andG3subtypeshavebeenobservedinimmunocompetent indi-viduals.PotentialriskfactorsforHEVtransmissionincludedpoor sanitation,persontopersoncontact,familywithmorethan4 mem-bers,parenteralbloodcontact,maletomalecontact,professional longexposurewithzoonoticreservoirsandrawandundercooked porkmeatandshellfish.
In order to assess the seroprevalence of hepatitis E, differ-entenzymeimmunoassays(EIA)have beenusedtodetectclass GandMimmunoglobulinsagainstHEV.Thecommercially avail-ableassayswerebasedontwomethodologies:ELISAandWestern Blot.Themajorityofstudies(22over27)usedoneormoreElisa assays[25–28,30–47],whilein5articleswereusedboth serolog-icalassaytypes[24,29,48–50].HEVRNAwasdetectedifsamples were positive for IgM and/or IgG in 12 of the selectedstudies [25,28,29,32,36,37,39,42,44,46,47,50].
Fig.1. Protocolutilizedtoselectarticlesforthereview.
Table1
Comparativesensitivityofeachcommercialassaybyregionsandstudypopulation.
Assay Regions Studypopulation(N) Anti-HEV HEV-RNA(%)
IgG(%) IgM(%) Assay-1 aCalabria GP(876) b17.8% – – aLombardy GP(2.489) b9.0% – Sardinia BD(402) 5.0% – – Sicilia GP(44) 4.7% – – Apulia GP(450) 2.7% 0.22% 0.22% BD(151) 1.3% – – Assay-2 Veneto GP(1.889) 2.6% RepublicofS.Marino GP(2.233) 1.5% – – Molise Ped(264) 0.4% – – Assay-3 Abruzzi BD(313) b49.0% 0.6% 0.6% Piedmont GP(73) 1.3% – – Assay-4 Tuscany BD(132) 9.1% – –
GP:generalpopulation;BD:blooddonors;Ped:pediatricstudy.
aDatafromthesamestudy[27].
Fig.2.MapofItalyshowingdifferentHEVseropositivityratesamongblooddonorsandgeneralpopulation..
Table1showsthereportedanti-HEVIgGseroprevalence accord-ingtotheassayusedintheItalianregions.Assay1(Dia.pro)was usedinthemajorityofstudiesandthereportedseroprevalence var-iedfrom17.8%inCalabriato2.7%and1.3%inApulia,respectively foundin generalpopulationand blood donors [27,29]. Assay-2 (Abbott)wasusedinthreeregionstodetecttheanti-HEVIgGin twostudiesamongthegeneralpopulation(seroprevalence rang-ingfrom2.6%and1.5%)andinapediatricstudy(seroprevalence 0.4%)inthreeregions[35,48,30].Thehighestandthelowest sero-prevalenceingeneralpopulationwerebothdetectedbyassay-3 (Wantai),respectively in Abruzzi (49%) and in Piedmont (1.3%) [25,28].BoththeAbruzziandApuliastudieshaveprovideddata onanti-HEVIgMandHEVRNAwithanoverallprevalenceranging from0.6%to0.22%,amongblooddonorsandgeneralpopulation, respectively.Intheformerstudythegenotypefoundwas3whilein thelatterstudywas1[25,29].ArecentstudyonItalianblooddonors hascomparedthediagnosticperformancesofassay-1and assay-3, demonstrating an overall concordance of 96%. Eventhought assay-3detectedaslightlylowerpositivityratethanassay-1,no significantdifferencein sensitivitywasobserved[47].However, ourBayesiananalysisofthereportedseroprevalencerates,among Italianregions,hasshownthatthescreeningmethodologywas ade-quateintermofspecificityonlyintwostudies.Thus,theestimated
HEVprevalenceinAbruzzi,CalabriaandLombardyreflectsthereal spreadingofinfection[25,27].
Discussion
Tothebestofourknowledge,thecurrentarticlerepresentsthe firstcriticalreviewof HEVIgGseroprevalenceinItaly. Thesera samplesof21.882individualswerecollectedfrom1978to2015 asreportedinthe27publications includedinthefinalanalysis. Theseroprevalenceranged from0.12%to49% withthehighest ratesbeinginthecentralregion.However,agreat geographical variabilitywasobservedamongtheItalianregionswithaaverage prevalenceof10,25%amongblooddonors.Thesefindingslaythe groundworkforthehypothesisthatdiversepredisposingfactors suchasdietaryhabits,environmentalcharacteristics,different dis-tributionsofzoonoticreservoirs,contaminatedsurfacewaters,the socio-economicandhygieniclevel,wouldsustainthespreadingof HEVinfectionalongtheItalianregionsinterdependently.
ThemajorityofautochthonouscasesofHEVinItalyseemtobe duetothezoonotictransmissionoftheinfectionfromdomestic andwildanimals.Todate,HEVinfectionshavebeendemonstrated indomesticpig,wildboars,rabbits,wilddearsandgoatsinItaly [51–55].Notsurprisingly,nearlyhalfofthewildboarsintheLatium
Table2
LaboratorydiagnosisofHEVinfection:Anti-HEVIgGAssays.
IgGassay Assaytype Antigenforcoating Strain Sensitivity Specificity
Abbott[91] – RecombinantORF-2andORF-3proteins Burmese 91% 96%
Adaltis(EIAgen)[92] Qualitativeindirect SyntheticantigensfromORF-2andORF-3 – 80.% 62.9% Dia.Pro[93] Qualitativeindirect 4syntheticpeptideswithconservative
epitopesofORF-2andORF-3Genotypes1, 2,3,and4
BurmeseandMexican 98% 96%
DSI[93] – RecombinantORF-2andORF-3peptides Genotype1,2and3
– 72% 99%
MPDiagnostics[95] – 3recombinantORF-2proteinsand 33-aminoacidsequencefromORF-3. Genotype2,3
– 98% 97%
Wantai[94] Qualitativeindirect RecombinantORF-2protein(PE2) Genotype1
Chinese 99,80% 99.9% W.Blot(Mikrogen)[93] Quantitativeindirect 4recombinantproteins(O2N,O2Mand
O2)fromORF-2andORF-3Genotype1,3
– 62% 99%
regionandin Tuscany(centralApenninesarea),had serological markersofHEVinfection[56,57].Asimilarhighseroprevalencewas demonstratedamongswineinSouthern(Calabria)andNorthern Italy but was remarkably low Piedmont region [58–61]. More-over,inNorthwesternItaly(LombardyandEmiliaRomagna),HEV contaminationwasdemonstratedinthemajorityofslurry sam-plesfrompigproductionfacilities[62].Inaddition,themajorityof indigenouscasesofacuteHEVwereclearlyrelatedtotheingestion ofraworundercookedlocalporkmeat[37].HEVcontamination hasbeenassessedinporkmeat-derivedproductsandinthe Ital-ianproductionchain.Interestingly,ahighnucleotidehomologyhas beenproveninhuman,swineandcontaminatedfoodsamplesfrom thesamegeographicalregionsand,asexpected,intermixedswine andhumangenomicsequenceswerefound[63–66].In fact,the inadequatecookingofcommerciallyavailableporkproductsdoes notinactivateeffectivelyHEVinfectivity[67].Inordertoprevent food-borneHEVinfection,consumingtheseproductscookedata temperatureof71◦Cforaminimumof20min,hasbeen exper-imentallyproven tobenecessary[68].Moreover,HEVhasbeen detectedinbaggedready-to-eat(RTE)vegetables,posingafurther concernregardingfoodsafetyandnewpotentialconsumers’risks [69].Genotype3appearstooccurinthemajorityofthelocally acquiredacuteHEVcases,bothinhumanandzoonoticreservoirs. However,genotype4hasbeenrecentlyreportedasanemerging indigenouspathogeninItalyaswellinFranceandGermany,and itseemsnolongerconfinedtoJapan,ChinaandSoutheasternAsia [70,71].Infact,asmalloutbreakwasreportedintheLatiumregion, in2011.Theisolatedstraindifferedgeneticallyfromtheidentified European4dand4fstrainsanditresembledthesubtype4dstrain isolatedinChinaamongtheswinepopulation[18].Furthermore,a genotype4strain,phylogenicallyrelatedtothehumanstrain iso-latedduringtheoutbreakincentralItaly,wasidentifiedinswine farmsinNorthernItalyandprovidedfurtherevidenceofaplausible cross-speciesinfectionandintroductionofanewHEVvariantina differentgeographicregion[72].
Althoughtheingestionofshellfishhasbeenreportedsince1980 asariskfactorinourcohortofpatientswithHEV,onlylatelyhas itsrolebeenassessedasanindicatorofmarinepollution[41].The bivalve molluscanshellfish sampleswere analyzedin indepen-dentstudiesinItaly,France,SpainandDenmarkfortheirability toconcentratetheviralparticlesfiltratedduringthefeeding pro-cess[73–76]. However, all the aforementioned studiesdid not supportthecontaminationofthemarineenvironment.No posi-tivesamplescollectedinthepotentiallycontaminatedsiteswere found.Thisresultcouldhavebeencausedbyeitheranundetectable quantityofviralparticlesorashort-livedenvironmental persis-tenceofHEV.Nevertheless,arecentstudyinShandongProvincein Chinaassessedtheseroprevalenceamong1028seafood-processing
workersof whom22.20% were anti-HEVIgG antibodypositive. Theincreaseinseroprevalencewasassociatedwithworking-time, thustoahigherlikelihoodtobeingexposedtocontaminatedraw seafood and semi-finished products[77]. Interestingly, time of exposurewastheonlyindependentvariablelinkedwithahigher anti-HEVprevalencefoundinourcohortofworkersatzoonoticrisk [28].
Thewaterbornerouteofinfectionhasbeenlargelyrecognized globally.Still,itsepidemiologicalimpactintheindustrialized coun-tries is unknown. However,HEV particles have been traced in theLatiumregionandintheTiberRiverthatrunsfromthe cen-tralApenninesregiontotheTyrrhenianSea,inItaly[78].Overall, thesefindingssuggestthatdifferentfactorscoulddeterminethe endemicityweobservedinItalyandthustheneedforfurther inves-tigation.
Arecentseroepidemiologicalstudycomparedagroupof res-identsintwodifferentregions:Lombardy(Abbiategrasso,Milan) andCalabria(Cittanova);reportingatwofoldincreasedHEV preva-lenceininthesouthernregion[27].Theauthorshaveexplained thedifferenceobserved,aslikelyconsequenceofthelowest socio-economicandhygienic/sanitaryconditionsinCalabria.Sinceonly theDia.proessay wasusedtodetermineHEVIgG positive, the resultmighttoreflecttherealspreadingofHEV,witha north-to-southgradient.Accordingtothisfinding,thehighestprevalence observedamongblooddonorsinAbruzziregionmightbea conse-quenceoftheinadequatesanitationandpoorhygienicconditions thatfollowedthedevastating earthquakethatstruckL’Aquilain 2009,causing over80000evacuatedfromtheirhomes,As mat-ter of thefact, an increase of enterictransmitted diseases was reportedsubsequentlytothecatastrophicenvironmentaland geo-logical changes [79].Moreover, thehighprevalenceratein the Abruzziregionislikelydueinparttothehighlysensitive Wan-taiassayused[80].BoththereportedIgMandRNAseroprevalence amongblooddonorsinAbruzziregionwas0.6%[25].
Furthermore,wefoundasimilarseroprevalencerateamong vol-unteerblooddonorsandthegeneralpopulation.Thisfindingought todenotethatthisspecificcohortcouldrepresenttheprevalence intheItalianpopulation,ingeneral[24,40].Moreover,weobserved that in themajority of cases HEVinfectionwas asymptomatic, anictericandself-limitingandanormalleveloftransaminaseshas beenalsoreported.Ontheotherhand,thisimpliesthatthe bio-chemicalandserologicalscreeningcurrentlyperformed;inorder toselectthehealthyblooddonorsmaybeunabletoidentityviremic donors.Indeed,viremicblooddonorswithanormalALTlevelhave beenreportedinGermanyandJapan[81,82].HEVRNAhasbeen detectedinblooddonationsandcasesoftransfusion-transmitted infectionhave beenreported worldwide [83–87].The
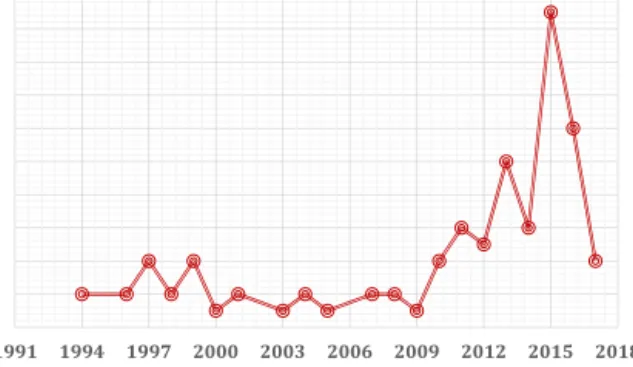
contami-Fig.3. NumberofHEVstudiesinItalyovertwoandone-halfdecades.
natedbloodcouldhaveanunder-recognizedroleasapotentialnew sourceofinfectionanditrequiresfurtherinvestigationsinItaly.
Wewereunabletoprovideaclearunderstandingofthe poten-tialimpact of theimmigration phenomenon among the Italian population since only two studies, in the same region, were includedin thefinal analysis [29,49]. However, according to a recent retrospective study on a small cohort of symptomatic migrants, hepatitis E appeared to be the main cause of acute viralhepatitis [88]. Nevertheless, 5%of 40 fecal samples,from asymptomaticimmigrantswerepositiveforHEVRNAsupportinga plausibleroleofimmigrantsas“symptom-freeHEVcarriers”[89]. Thecurrentstudysuggeststheexistenceofagreatvariability intheseroprevalenceofHEVinItaly.Thisresultcouldbepartially explainedbytheheterogeneityinthesensitivityandspecificityof theimmunoassaysused,bysmallnumberofstudiesincludedas wellasthenumberofsamplestested;andthenarrowwindowof collectingsamplesperiod.Allsamplesfromthegeneralpopulation andtheblooddonorshavebeenanalyzedwithfourcommercially availableELISAtest:Dia.pro,Abbott,Wantai,Adaltis.Performance comparisonamongdifferentassayswasreportedbyarecentstudy, whichshowedaverygoodconcordancebetweentheDia.proand Wantaiassay[47].
TheglobalburdenofhepatitisEisstillunderestimateddueto thesub-optimalcommercialassaysavailable[90].
Infact,serologicalassaysbasedondifferentgenotypes,using recombinantproteinsorsyntheticpeptides,varygreatlyinterm ofsensitivityandspecificity(Table2)[91–95].Moreover,theassay sensitivityishigherinsymptomaticcasesthanintheasymptomatic ones[91].Nevertheless,thespecificityofthescreening methodol-ogy,toobtainvalidvalueofHEVprevalence,differaccordingto theinfectionendemicity.However, theselimitationsempathize thenecessityofa comparablystandardseroprevalencestudyat anationallevel,inordertoestimatetherealprevalenceof Hep-atitisEandtocreateaninterventionalplandirectedataregional level.Clearly,differentdietaryhabitscan’t alonedeterminethe variabilityobservedinourstudies.
Conclusion
TheWorldHealthOrganizationdefinesasanemerging zoono-sisanydiseasethatis“newlyrecognizedornewlyevolved,orhas shownanincreaseinincidenceorexpansioningeographical,host orvectorrange”[96].Wedo notknowwhetherHEVistrulyan emerginginfectionorwhetheritisduetoanincreasedawareness andunderstandingofHEVintheWesterncountries(Fig.3).
Althoughaphylogeneticandevolutionaryanalysishasstated thatHEVmighthavebeenpresentintheItalianterritorysincethe early90s,nowadaysitremains asilentand understudiedentity [66].Atthepresent,HEVisundoubtedlyendemicinItaly. How-ever,thelackofcommerciallyapproveddiagnosticassays[95],the
nizedinfectiousdiseases.HEVshouldalwaysbeconsideredinthe differentialdiagnosisofacuteviralhepatitis.
Funding Nofundingsources. Competinginterests Nonedeclared. Ethicalapproval Notrequired. Acknowledgments
PartiallyfundedbytheUlyssesGrant(BorsadistudioUlisse) oftheUniversityofSassariSchoolofMedicine,andbytheJohns HopkinsPediatricLiverCenter.
References
[1]ReinDB,StevensGA,TheakerJ,WittenbornJS,WiersmaST.Theglobalburden ofhepatitisEvirusgenotypes1and2in2005.Hepatology2012;55(4):988–97.
[2]LozanoR,NaghaviM,ForemanK,LimS,ShibuyaK,AboyansV,etal.Global andregionalmortalityfrom235causesofdeathfor20agegroupsin1990and 2010:asystematicanalysisfortheGlobalBurdenofDiseaseStudy2010.Lancet 2012;380:2095–128.
[3]KhurooMS.Studyofanepidemicofnon-A,non-Bhepatitis:possibilityof anotherhumanhepatitisvirusdistinctfrompost-transfusionnon-A,non-B type.AmJMed1980;68:818–23.
[4]YamashitaT,MoriY,MiyazakiN,ChengRH,YoshimuraM,UnnoH,etal. Biolog-icalandimmunologicalcharacteristicsofhepatitisEvirus-likeparticlesbased onthecrystalstructure.ProcNatlAcadSciUSA2009;106(31):12986–91.
[5]SmithDB,PurdyMA,SimmondsP.Geneticvariabilityandtheclassificationof hepatitisEvirus.JVirol2013;87(April(8)):4161–9.
[6]Lu L, Li C, Hagedorn CH. Phylogenetic analysis of global hepatitis E virussequences:genetic diversity,subtypesandzoonosis. RevMed Virol 2006;16:5–36.
[7]GuthmannJP1,KlovstadH,BocciaD,HamidN,PinogesL,NizouJY,etal.Alarge outbreakofhepatitisEamongadisplacedpopulationinDarfur,Sudan,2004: theroleofwatertreatmentmethods.ClinInfectDis2006;42:1685–91.
[8]NicandE1,ArmstrongGL,EnoufV,GuthmannJP,GuerinJP,CaronM,etal. GeneticheterogeneityofhepatitisEvirusinDarfur,Sudan,andneighboring Chad.JMedVirol2005;77(4):519–21.
[9]AwsathiS,RawatV,RawatCM,SemwalV,BartwalSJ.Epidemiological inves-tigationofthejaundiceoutbreakinLalkuan,Nainitaldistrict,Uttarakhand. IndianJCommunityMed2014;39:94–7.
[10]Sedyaningsih-MamahitER,LarasaRP,LarasK,SidemenA,SukriN,Sabaruddin N,etal.FirstdocumentedoutbreakofhepatitisEvirustransmissioninJava, Indonesia.TransRSocTropMedHyg2002;96:398–404.
[11]VillalbaMDLCM,LayLDLAR,ChandraV,CorredorMB,FrometaSS,MorenoAG, etal.HepatitisEvirusgenotype1,Cuba.EmergInfectDis2008;14:1320–2.
[12]Al-NasrawiKK,Al-DiwanJK,Al-HadithiTS,SalehAM.ViralhepatitisEoutbreak inAl-Sadrcity,Baghdad,Iraq.EastMediterrHealthJ2010;16:1128–32.
[13]KhurooMS,TeliMR,SkidmoreS,SofiMA,KhurooMI.Incidenceandseverityof viralhepatitisinpregnancy.AmJMed1981;70:252–5.
[14]WilhelmB,MuellnerP,PearlDL,Raji ´cA3,HoudeA,McEwenSA.Preliminary molecularepidemiologicalinvestigationofhepatitisEvirussequencesfrom Quèbec,Canada.PrevVetMed2015;118(4):359–69.
[15]Amon JJ,DrobeniucJ, BowerWA, Maga ˜naJC, EscobedoMA,WilliamsIT, etal.LocallyacquiredhepatitisEvirusinfection,ElPaso,Texas.JMedVirol 2006;78(6):741–6.
[16]YapaCM,FurlongC,RosewellA,WardKA,AdamsonS,ShadboltC,etal.First reportedoutbreakoflocallyacquiredhepatitisEvirusinfectioninAustralia. MedJ2016;204(7):274.
[17]TesséS,LioureB,ForneckerL,WendlingMJ,Stoll-KellerF,BigaillonC,etal. Cir-culationofgenotype4hepatitisEvirusinEurope:firstautochthonoushepatitis EinfectioninFrance.JClinVirol2012;54:197–200.
[18]GarbugliaAR,ScognamiglioP,PetrosilloN,MastroianniCM,SordilloP,Gentile D,etall.HepatitisEvirusgenotypes4outbreak,Italy,2011.EmergInfectDis 2013;19(1):110–4.
[19]Yazaki Y,SugawaraK,HondaM,OhnishiH,NagashimaS,Takahasmi M, etal.Characteristicsof20patientswithautochthonousacutehepatitisEin
Hokkaido,Japan:firstreportofbilateralfacialpalsyfollowingtheinfection withgenotype4hepatitisEVirus.TohokuJExpMed2015;236(4):263–71.
[20]TakahashiM,NishizawaT,NagashimaS,JirintaiS,KawakamiM,SonodaY,etal. MolecularcharacterizationofanovelhepatitisEvirus(HEV)strainobtained fromawildboarinJapanthatishighlydivergentfromthepreviously recog-nizedHEVstrains.VirusRes2014;180:59–69.
[21]SridharS,TengJLL,ChiuTH,LauSKP,WooPCY.HepatitisEvirusgenotypesand evolution:emergenceofcamelhepatitisEvariants.IntJMolSci2017;18(4).
[22]HartlJ,OttoB,MaddenRG,WoolsonKL,KristonL,etal.HepatitisE seropreva-lenceinEurope:ameta-analysis.Viruses2016;8(8):211.
[23]ISTAT.DemographicIndicators.Italy,Rome;2016.www.istat.it/en/archive/ demographic+indicators.[Accessed3September2016].
[24]ZanettiAR,DawsonGJ.HepatitistypeEinItaly:aseroepidemiologicalsurvey: studygroupofhepatitisE.JMedVirol1994;42(3):318–20.
[25]LucarelliC,SpadaE,TalianiG,ChionneP,MadonnaE,MarcantonioC,etal.High prevalenceofanti-hepatitisEvirusantibodiesamongblooddonorsincentral Italy,FebruarytoMarch2014.EuroSurveill2016;21(30).
[26]PuttiniC,RiccioML,RediD,TordiniG,CeneriniM,RomanelloF,etal. Sero-prevalenceofhepatitisEvirus(HEV)infectioninblooddonorsandrenal transplant recipients:a retrospectivestudy from centralItaly.InfezMed 2015;23(3):253–6.
[27]ZuinM,CasertaC,RomanòL,MeleA,ZanettiA,CannatelliR,GiorginiA,etal. SeroepidemiologyofHEVandHAVintwopopulationswithdifferent socio-economiclevelsandhygienic/sanitaryconditions.EurJClinMicrobiolInfect Dis2017;36(3):479–85.
[28]CarusoC,PelettoS,RosamiliaA,ModestoP,ChiavacciL,SonaB,etal.Hepatitis Evirus:across-sectionalserologicalandvirologicalstudyinpigsandhumans atzoonoticriskwithinahigh-densitypigfarmingarea.TransboundEmergDis 2016,http://dx.doi.org/10.1111/tbed.1253.
[29]ScottoG,MartinelliD,CentraM,QuerquesM,VittorioF,etal.Epidemiological andclinicalfeaturesofHEVinfection:asurveyinthedistrictofFoggia(Apulia, SouthernItaly).EpidemiolInfect2013;142(2):287–94.
[30]RipabelliG,SammarcoML,CampoT,MontanaroC,D’AscenzoE,GrassoGM. Prevalenceofantibodiesagainstentericallytransmittedviralhepatitis(HAV andHEV)amongadolescentsinaninlandterritoryofcentralItaly.EurJ Epi-demiol1997;13(1):45–7.
[31]RomanòL,PaladiniS,TagliacarneC,CanutiM,BianchiS,ZanettiAR.Hepatitis EinItaly:along-termprospectivestudy.JHepatol2011;54(1):34–40.
[32]Zanetti AR,SchlauderGG,RomanòL,TanziE,FabrisP,DawsonGJ,etal. Identification ofa novel variant ofhepatitis E virusin Italy. Med Virol 1999;57(4):356–60.
[33]FabriziF,LunghiG,BacchiniG,CortiM,PaganoA,LocatelliF.HepatitisEvirus infectioninhaemodialysispatients:aseroepidemiologicalsurvey.NephrolDial Transplant1997;12(1):133–6.
[34]TabibiR,BaccaliniR,BarassiA,BonizziL,BrambillaG,ConsonniD,etal. Occu-pationalexposuretozoonoticagentsamongagriculturalworkersinLombardy Region,northernItaly.AnnAgricEnvironMed2013;20(4):676–81.
[35]GessoniG1,ManoniF.HepatitisEvirusinfectioninnorth-eastItaly: sero-logical study in the open population and groups at risk. J Viral Hepat 1996;3(4):197–202.
[36]Stroffolini T,Rapicetta M,ChionneP, EsvanR, MadonnaE, Lombardo F, etal.Evidenceforthepresenceofautochthonous(locallyacquired)cases ofacutehepatitisEvirusinfectionsinItalysincethe80s.EurJInternMed 2015;26(5):348–50.
[37]TarantinoG,BagnarelliP,MarzioniM,MarinelliK,SuraceG,TrainiS,etal. HepatitisEinaregionofItaly:anemergingautochthonousinfection?DigLiver Dis2016;48(11):1340–5.
[38]LaniniS,GarbugliaAR,LapaD,PuroV,NavarraA,PergolaC.Epidemiology ofHEVintheMediterraneanbasin:10-yearprevalenceinItaly.BMJOpen 2015;5(7):e007110.
[39]La Rosa G, Muscillo M,VennarucciVS, Garbuglia AR, La Scala P, Capo-bianchiMR.HepatitisEvirusinItaly:molecularanalysisoftravel-relatedand autochthonouscases.JGenVirol2011;92(Pt.7):1617–26.
[40]PaviaM,IiritanoE,VerattiMA,AngelilloIF.PrevalenceofhepatitisEantibodies inhealthypersonsinsouthernItaly.Infection1998;26(1):32–5.
[41]CacopardoB,RussoR,PreiserW,BenantiF,BrancatiG,NunnariA.Acute hepati-tisEinCatania(easternSicily)1980–1994:theroleofhepatitisEvirus.Infection 1997;25(5):313–6.
[42]CacciolaI,MessineoF,CacopardoB,DiMarcoV,GalliC,SquadritoG,etal. HepatitisEvirusinfectionasacauseofacutehepatitisinSouthernItaly.Dig LiverDis2011;43(12):996–1000.
[43]LaFauciV,FacciolàA,RisoR,CalimeriS,LoGiudiceD,SqueriR.Seroprevalence ofhevantibodiesinasampleofpregnantwomeninthecityofMessina.Ann Ig2017;29(3):232–8.
[44]MasiaG,OrrùG,LiciardiM,DesogusG,CoppolaRC,MurruV,etal.Evidence ofhepatitisEvirus(HEV)infectioninhumanandpigsinSardinia,Italy.JPrev MedHyg2009;50(4):227–31.
[45]RapicettaM,MonarcaR,KondiliLA,ChionneP,MadonnaE,MadedduG,etal. HepatitisEvirusandhepatitisAvirusexposuresinanapparentlyhealthy high-riskpopulationinItaly.Infection2013;41(1):69–76.
[46]CandidoA,TaffonS,ChionneP,PisaniG,MadonnaE,DettoriS,etal.Diagnosis ofHEVinfectionbyserologicalandreal-timePCRassays:astudyonacute non-A–Chepatitiscollectedfrom2004to2010inItaly.BMCResNotes2012;5:297.
[47]RiccoG,BoninoF,LanzaM,ScatenaF,AlfieriCM,MessaP,etal.New immunoas-saysfortotal,IgAandIgMantibodiesagainsthepatitisEvirus:prevalencein
Italianblooddonorsandpatientswithchronicliverorkidneydiseases.Dig LiverDis2016;48(5):536–41.
[48]RapicettaM,KondiliLA,PretolaniS,StroffoliniT,ChionneP,VillanoU,etal. Seroprevalenceandanti-HEVpersistenceinthegeneralpopulationofthe RepublicofSanMarino.JMedVirol1999;58(1):49–53.
[49]DeDonnoA,ChironnaM,CracaR,PaianoA,ZizzaA,etal.Anti-HEV seropreva-lenceintheareaofLecce.AnnIg2003;15(3):199–205.
[50]DeSabatoL,DiBartoloI,MontomoliE,TrombettaC,RuggeriFM,Ostanello F.RetrospectivestudyevaluatingseroprevalenceofhepatitisEvirusinblood donorsandinswineveterinariansinItaly(2004).ZoonosesPublicHealth 2017;64(4):308–12.
[51]MoniniM,DiBartoloI,IaniroG,AngeloniG,MagistraliCF,OstanelloF,etal. Detectionandmolecularcharacterizationofzoonoticvirusesinswinefecal samplesinItalianpigherds.ArchVirol2015;160(10):2547–56.
[52]MartelliF,CaprioliA,ZengariniM,MarataA,FiegnaC,DiBartoloI,etal. Detec-tionofhepatitisEvirus(HEV)inademographicmanagedwildboar(Susscrofa scrofa)populationinItaly.VetMicrobiol2008;126(1–3):74–81.
[53]DiBartoloI,DeSabatoL,MarataA,MartinelliN,MagistraliCF,MoniniM,etal. SerologicalsurveyofhepatitisEvirusinfectioninfarmedandpetrabbitsin Italy.ArchVirol2016;161(5):1343–6.
[54]DiBartoloI,PonterioE,AngeloniG,MorandiF,OstanelloF,NicolosoS,etal. PresenceofhepatitisEvirusinaREDdeer(Cervuselaphus)populationinCentral Italy.TransboundEmergDis2017;64:137–43.
[55]DiMartinoB,DiProfioF,MelegariI,SarcheseV,RobettoS,MarsilioF,etal. DetectionofhepatitisEvirus(HEV)ingoats.VirusRes2016;225:69–72.
[56]MontagnaroS,DeMartinisC,SassoS,CiarciaR,DamianoS,AulettaL,etal.Viral andantibodyprevalenceofhepatitisEinEuropeanwildboars(Susscrofa)and huntersatzoonoticriskintheLatiumregion.JCompPathol2015;153(1):1–8.
[57]MazzeiM,NardiniR,VerinR,ForzanM,PoliA,TolariF.Serologicand molec-ularsurveyforhepatitisEvirusinwildboar(Susscrofa)inCentralItaly.New MicrobesNewInfect2015;7:41–7.
[58]SerraccaL,BattistiniR,RossiniI,MignoneW,PelettoS,BoinC,etal.Molecular investigationonthepresenceofhepatitisEvirus(HEV)inwildgamein North-WesternItaly.FoodEnvironVirol2015;7(3):206–12.
[59]CarusoC,ModestoP,BertoliniS,PelettoS,AcutisPL,DondoA,etal. Sero-logicalandvirologicalsurveyofhepatitisEvirusinwildboarpopulations innorthwestern Italy: detectionof HEV subtypes3e and 3f. Arch Virol 2015;160(January(1)):153–60.
[60]CostanzoN,SarnoE,PerettiV,CiambroneL,CasalinuovoF,SantoroA. Sero-logicalandmolecularinvestigationofswinehepatitisEvirusinpigsraisedin SouthernItaly.JFoodProt2015;78(11):2099–102.
[61]DiBartoloI,PonterioE,CastelliniL,OstanelloF,RuggeriFM.Viraland anti-bodyHEVprevalence inswine at slaughterhousein Italy.VetMicrobiol 2011;149(3–4):330–8.
[62]LaRosaG,DellaLiberaS,BrambillaM,BisagliaC,PisaniG,CiccaglioneAR,etal. HepatitisEvirus(genotype3)inslurrysamplesfromswinefarmingactivities inItaly.FoodEnvironVirol2016;9(2):219–29.
[63]DiBartoloI,Diez-ValcarceM,VasickovaP,KralikP,HernandezM,AngeloniG, etal.HepatitisEvirusinporkproductionchaininCzechRepublic,Italy,and Spain,2010.EmergInfectDis2012;18(8):1282–9.
[64]DiBartoloI,AngeloniG,PonterioE,OstanelloF,RuggeriFM.Detectionof hep-atitisEvirusinporkliversausages.IntJFoodMicrobiol2015;193:29–33.
[65]GarbugliaAR,AlessandriniAI,PavioN,TesséS,GrignoloS,ViscoliC,etal.Male patientwithacutehepatitisEinGenoa,Italy:figatelli(porkliversausage)as probablesourceoftheinfection.ClinMicrobiolInfect2015;21(1):e4–6.
[66]MontesanoC,GiovanettiM,CiottiM,CellaE,LoPrestiA,GrifoniA,etal. Hep-atitisEviruscirculationinItaly:phylogeneticandevolutionaryanalysis.Hepat Mon2016;16(3):e31951.
[67]FeaginsAR,OpriessnigT,GuenetteDK,HalburPG,MengXJ.Inactivationof infectioushepatitisEviruspresentincommercialpigliverssoldinlocalgrocery storesintheUnitedStates.IntJFoodMicrobiol2008;123(1–2):32–7.
[68]BarnaudE,RogéeS,GarryP,RoseN,PavioN.Thermalinactivationofinfectious hepatitisEvirusinexperimentallycontaminatedfood.ApplEnvironMicrobiol 2012;78(15):5153–9.
[69]TerioV,BottaroM,PavoniE,LosioMN,SerrainoA,GiacomettiF,etal. Occur-renceofhepatitisAandEandnorovirusGIandGIIinready-to-eatvegetables inItaly.IntJFoodMicrobiol2017;249:61–5.
[70]BouamraY,GérolamiR,ArzouniJP,GrimaudJC,LafforgueP,NelliM,etal. EmergenceofautochthonousinfectionswithhepatitisEvirusofgenotype4in Europe.Intervirology2014;57(1):43–8.
[71]WichmannO,SchimanskiS,KochJ,KohlerM,RotheC,PlentzA,etal. Phyloge-neticandcase-controlstudyonhepatitisEvirusinfectioninGermany.JInfect Dis2008;198(12):1732–41.
[72]MonneI,CeglieL,MartinoDI,NataleG,ZamprognaS,MorrealeA,etal. HepatitisEvirus genotype4in apig farm,Italy,2013. EpidemiolInfect 2015;143(3):529–33.
[73]IaconelliM,PurpariG,DellaLiberaS,PetriccaS,GuercioA,CiccaglioneA,etal. HepatitisAandEvirusesinwastewaters,inriverwaters,andinbivalve mol-luscsinItaly.FoodEnvironVirol2015;7(4):316–24.
[74]GrodzkiM,SchaefferJ,PiquetJC,LeSauxJC,ChevéJ,OllivierJ,etal. Bioac-cumulationefficiency,tissuedistribution,andenvironmentaloccurrenceof hepatitisEvirusinbivalveshellfishfrom France.Appl EnvironMicrobiol 2014;80(14):4269–76.
[75]MesquitaJR,OliveiraD,RivadullaE,Abreu-SilvaJ,VarelaMF,RomaldeJL,etal. HepatitisEvirusgenotype3inmussels(Mytilusgalloprovinciallis)Spain.Food Microbiol2016;58:13–5.
andrelatedriskfactorsamongseafoodprocessingworkers:across-sectional surveyinShandongProvince,China.IntJInfectDis2016;49:62–6.
[78]MarcheggianiS,D’UgoE,PuccinelliC,GiuseppettiR,D’AngeloAM,GualerziCO, etal.Detectionofemergingandre-emergingpathogensinsurfacewatersclose toanurbanarea.IntJEnvironResPublicHealth2015;12(5):5505–27.
[79]NigroG,BottoneG,MaioraniD,TrombatoreF,FalascaS,BrunoG.Pediatric epidemicofSalmonellaentericaSerovarTyphimuriumintheareaofL’Aquila, Italy,fouryearsafteracatastrophicearthquake.IntJEnvironResPublicHealth 2016;13:475.
[80]KodaniM,KamiliNA,Tejada-StropA,PoeA,DennistonMM,DrobeniucJ, etal.VariabilityintheperformancecharacteristicsofIgGanti-HEVassays anditsimpactonreliabilityofseroprevalenceratesofhepatitisE.JMedVirol 2017;89(June(6)):1055–61.
[81]Vollmer T,Diekmann J, EberhardtM,Knabbe C, Dreier J. HepatitisEin blooddonors:investigationofthenaturalcourseofasymptomaticinfection, Germany,2011.EuroSurveill2016;21(35).
[82]FukudaS,SunagaJ,SaitoN,FujimuraK,ItohY,SasakiM,etal.Prevalenceof antibodiestohepatitisEvirusamongJapaneseblooddonors:identificationof threeblooddonorsinfectedwithagenotype3hepatitisEvirus.JMedVirol 2004;73(4):554–61.
[83]ShresthaAC,FlowerRLP,SeedCR,KellerAJ,HarleyR,ChanH-T,etal.Hepatitis EvirusRNAinAustralianblooddonations.Transfusion2016;56:3086–93.
[84]TedderRS,TettmarKI,BrailsfordSR1,SaidB,Ushiro-LumbI,KitchenA,etal. Virology,serology:anddemographyofhepatitisEviremicblooddonorsin SouthEastEngland.Transfusion2016;56(6Pt2):1529–36.
[85]FuseK,MatsuyamaY,MoriyamaM,MiyakoshiS,ShibasakiY,TakizawaJ,etal. Lateonsetpost-transfusionhepatitisEdevelopingduringchemotherapyfor acutepromyelocyticleukemia.InternMed2015;54(6):657–61.
[86]MatsuiT,KangJH,MatsubayashiK,YamazakiH,NagaiK,SakataH,etal.Rare caseoftransfusion-transmittedhepatitisEfromthebloodofadonorinfected withthehepatitisEvirusgenotype3indigenoustoJapan:viraldynamicsfrom onsettorecovery.HepatolRes2015;45(6):698–704.
[88]BradaniniL,YoukeeD,FabrisP,RomanòL,BrunettiE,GiordaniMT.Acute hepatitisEvirusinfectioninamigrantpopulationinNorthEastItaly:A ret-rospectiveanalysis.TravelMedInfectDis2017;0(0).
[89]IdoloA,SerioF,LugoliF,GrassiT,BagordoF,GuidoM,etal.Identification ofHEVinsymptom-freemigrantsandenvironmentalsamplesinItaly.JViral Hepat2013;20(6):438–43.
[90]Kmush BL, Labrique AB,Dalton HR,Ahmed ZB,Ticehurst JR,etal. Two generationsofgoldstandards:theimpactofadecadeinhepatitisEvirus testing innovation on population seroprevalence. Am J Trop Med Hyg 2015;93(4):714–7.
[91]MyintKSA,EndyTP,GibbonsRV,LarasK,MammenJrMP,SeyaningsihER,etal. EvaluationofdiagnosticassaysforhepatitisEvirusinoutbreaksettings.JClin Microbiol2006;44(4):1581–3.
[92]WuWC,SuCW,YangJY,LinSF,ChenJY,WuJC.Applicationofserologicassays fordiagnosingacutehepatitisEinnationalsurveillanceofanonendemicarea. JMedVirol2014;86(4):720–8.
[93]NorderH,KarlssonM,ÅMellgren,KonarJ,SandbergE,LassonA,CastedalM, MagniusL,LaggingM.Diagnosticperformanceoffiveassaysforanti-hepatitis EvirusIgGandIgMinalargecohortstudy.JClinMicrobiol2016;54:549–55.
[94]WantaiHEV-IgGELISA,BeijingWantaiBiologicalPharmacyEnterpriseCo., Ltd.; 2017. http://www.ystwt.cn/Presentation/Wantai%20HEV%20ELISA.pdf. [Accessed14June2017].
[95]Shrestha AC,FlowerRL, SeedCR, StramerSL,FaddyHM.Acomparative studyofassayperformanceofcommercialhepatitisEvirusenzyme-linked immunosorbentassaykitsinAustralianblooddonorsamples.JBloodTransfus 2016;2016:9647675.
[96]WorldHealthOrganization.EmergingZoonoses.WHO;2016.http://www.who. int/zoonoses/emergingzoonoses/en/.[Accessed10October2016].